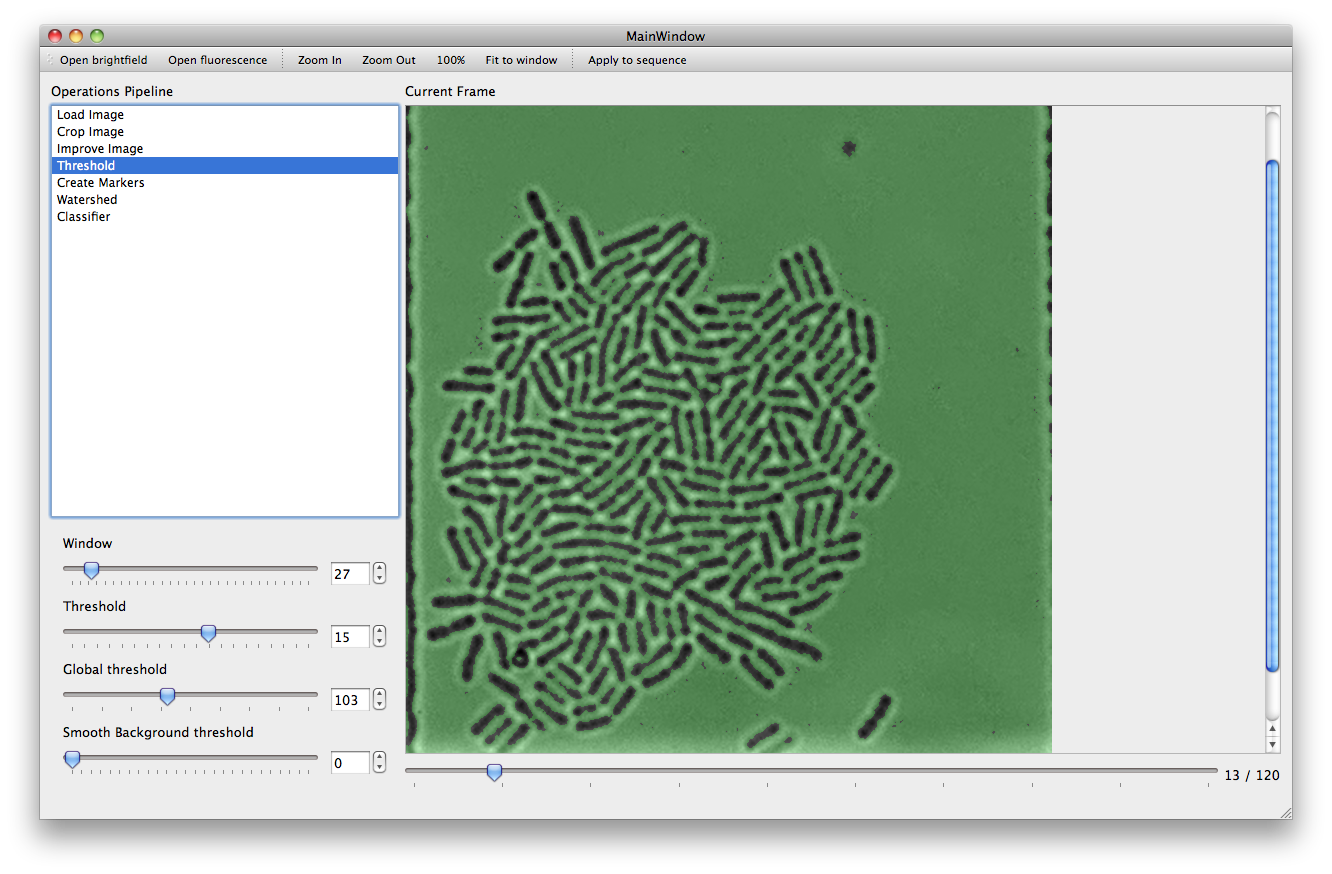
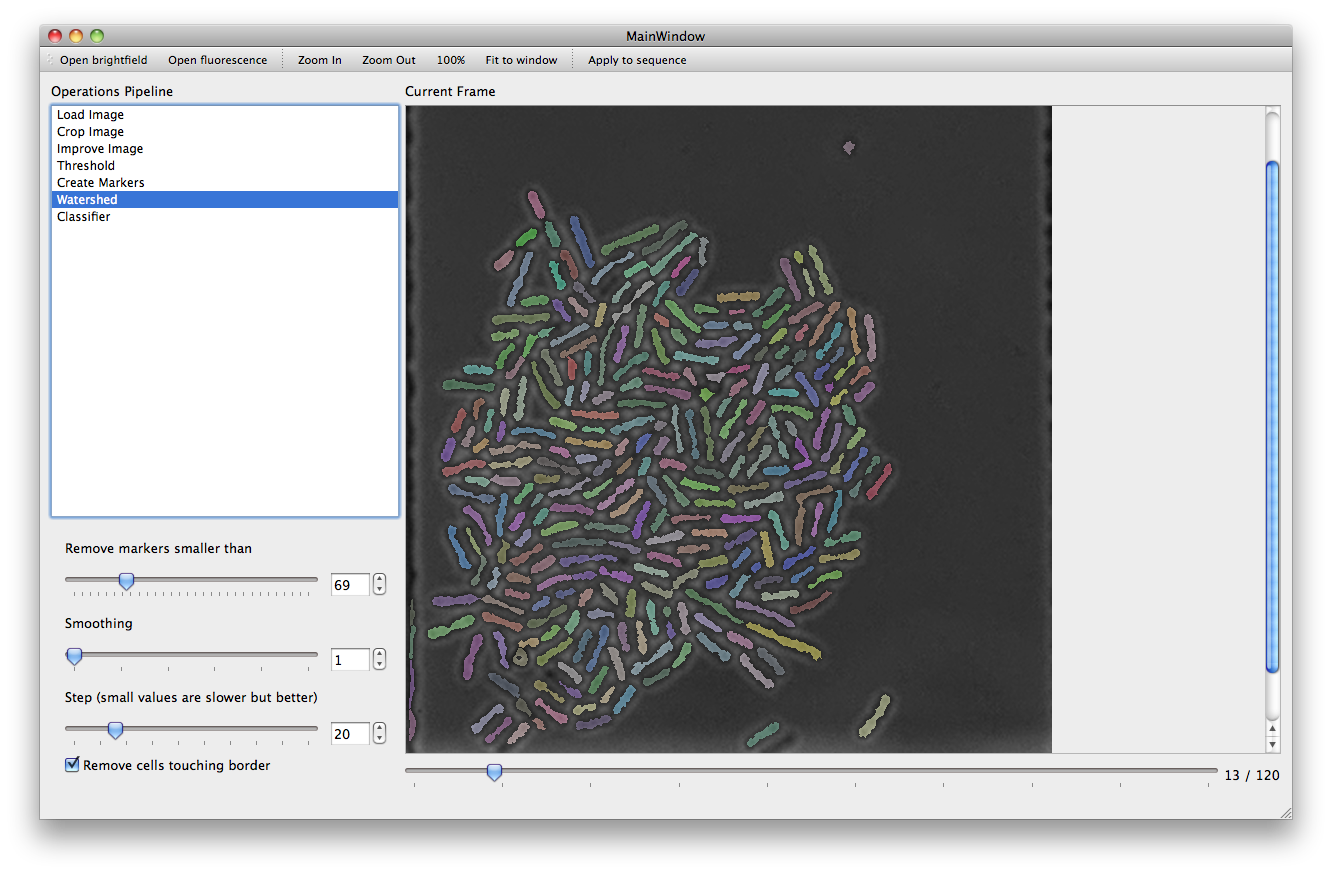
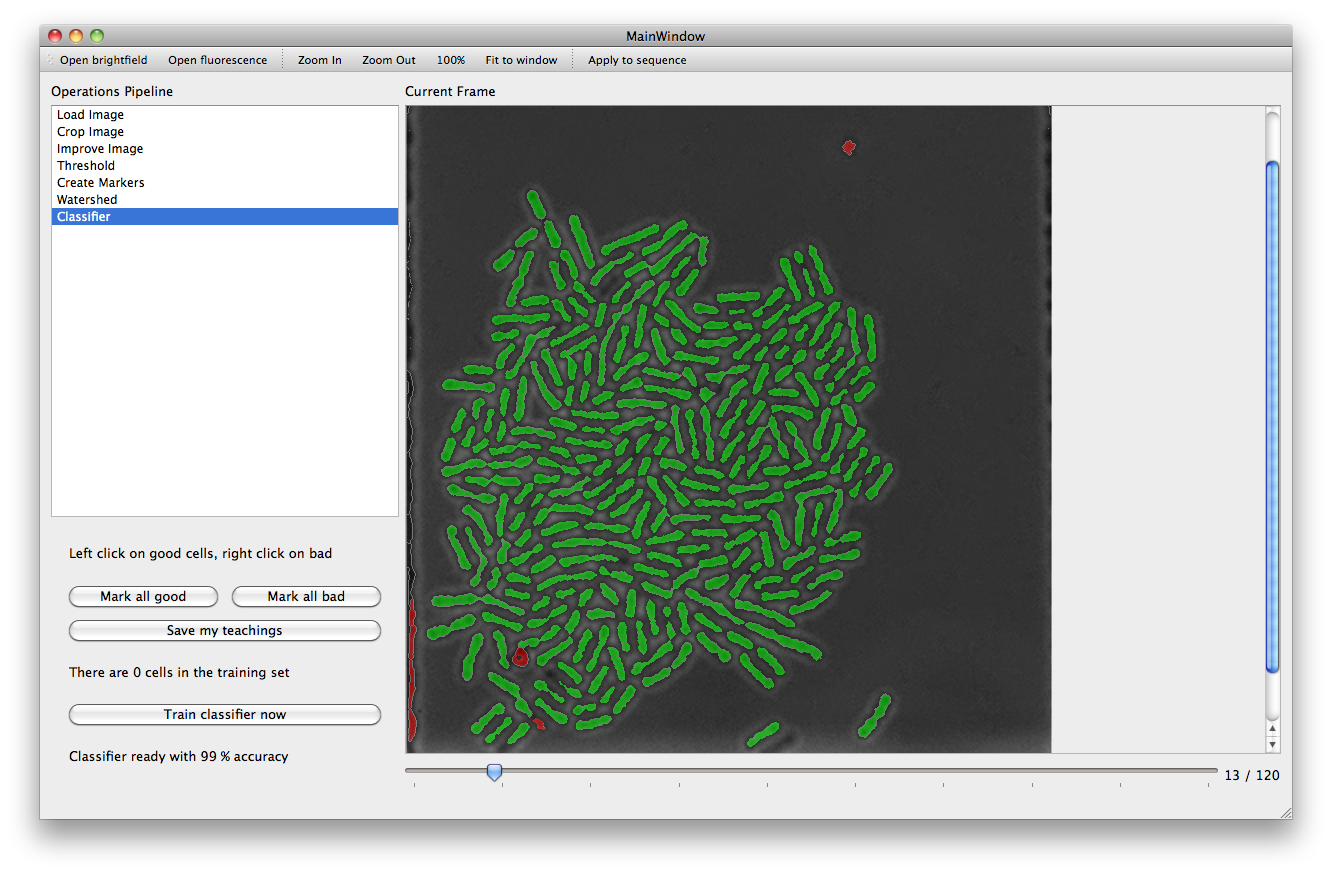
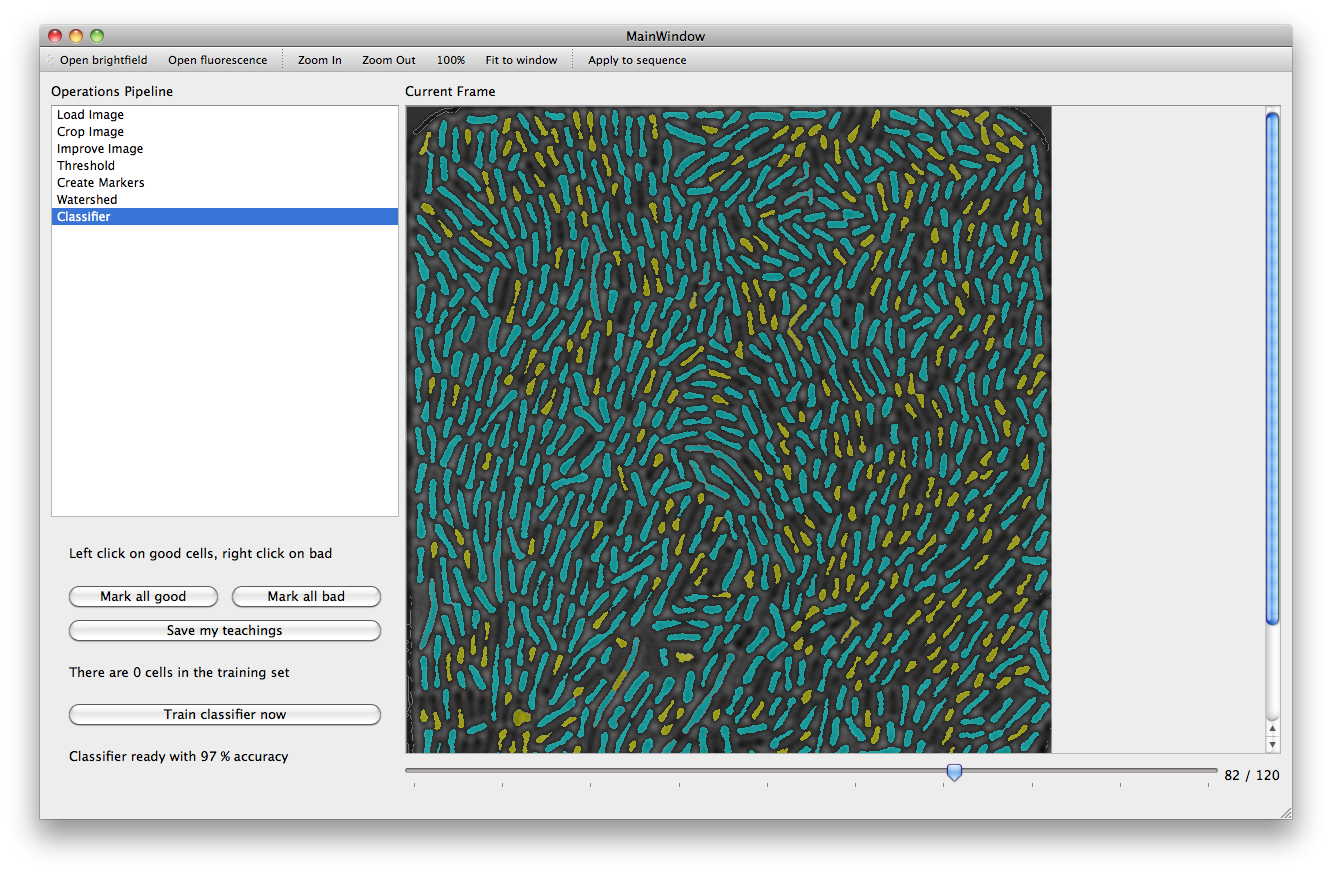
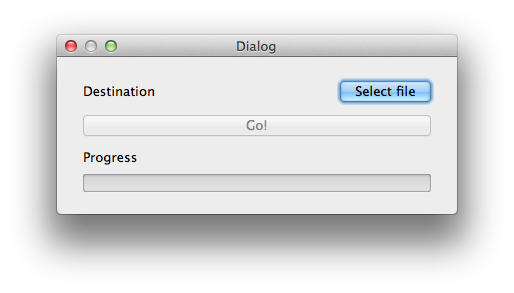
With the Threshold action a set of filters are applied to the image in order to mark areas as background. The more pixels are marked as background, the better the performance of subsequent steps will be. However caution must be taken to ensure that the chosen parameters are not over optimized for the current frame leaving other frames with a poor background (it is possible to use the slider under the main picture frame to look at later frames).
The window and threshold parameters control the adaptive threshold algorithm, which marks a pixel as background if its average value is lower than that of its neighbors within a window of the given size. The global threshold parameter controls a simple brightness filter: any pixel under that brightness is marked as background. Smooth background controls an adaptive filter which might mark certain small objects as background.